ARC: Assessment of Interface Residue Conformation using an Ensemble of Graph Neural Networks
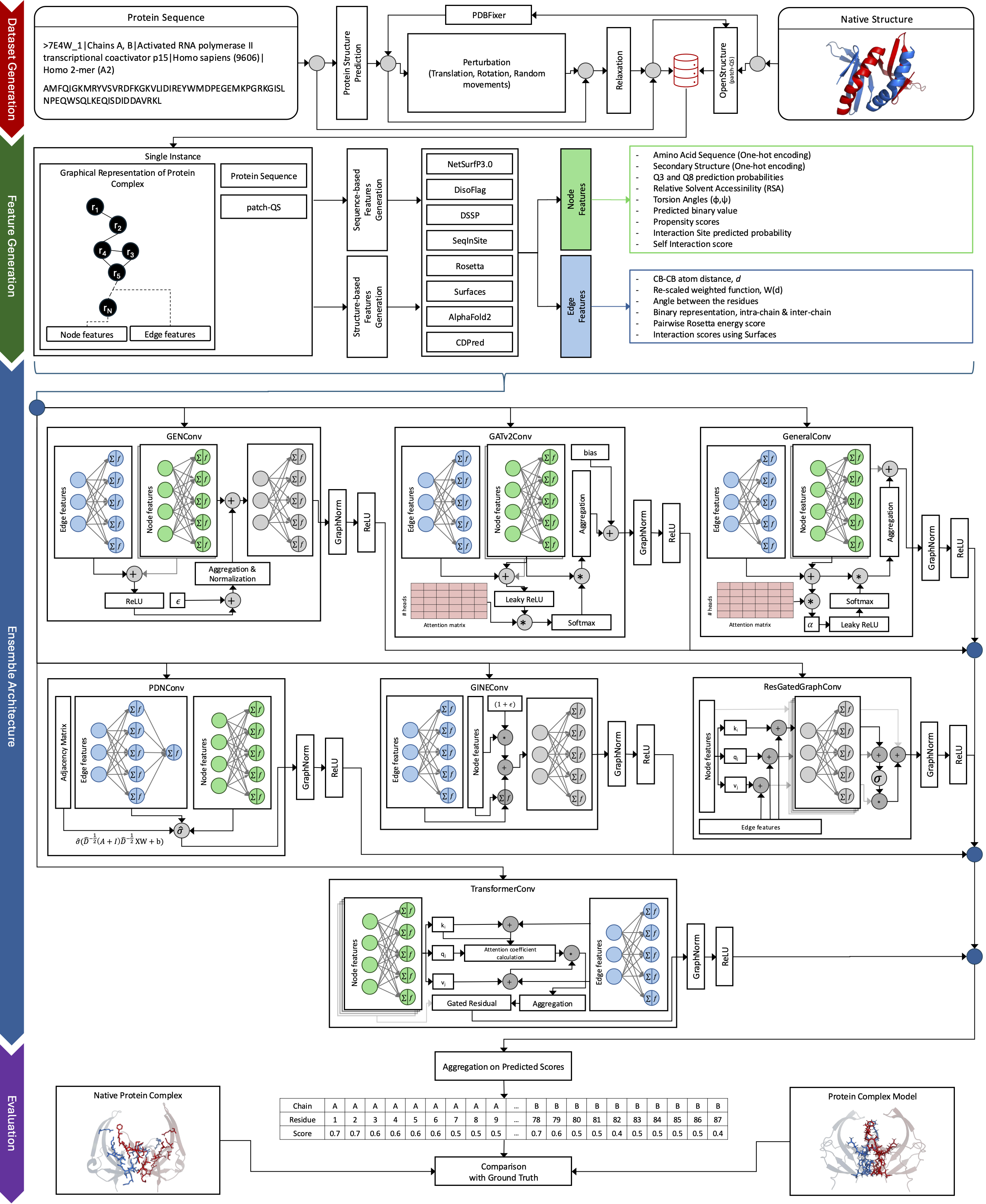
ARC is a single-model quality assessment method for protein complexes that assigns per-residue scores on modeled chain-chain interfaces using an ensemble of graph neural networks. The method is designed to capture local interface geometry and physicochemical context, enabling accurate identification and discrimination of true interface residues.
By focusing on residue-level predictions, ARC complements global assembly-level metrics, which may remain high despite inaccuracies in interface placement. ARC therefore provides a more direct signal of interface correctness, particularly in structurally complex or higher-order assemblies.
Download
The source code and evaluation pipeline are available on GitHub: https://github.com/zwang-bioinformatics/ARC/
Reference
Shrestha, B., Siciliano, A. J., Huang, G., Bao, Y., and Wang, Z.
ARC: Assessment of Interface Residue Conformation using an Ensemble of Graph Neural Networks.
Proteins: Structure, Function, and Bioinformatics, 2026.
(submitted)
Contact
Dr. Zheng Wang
Department of Computer Science
University of Miami